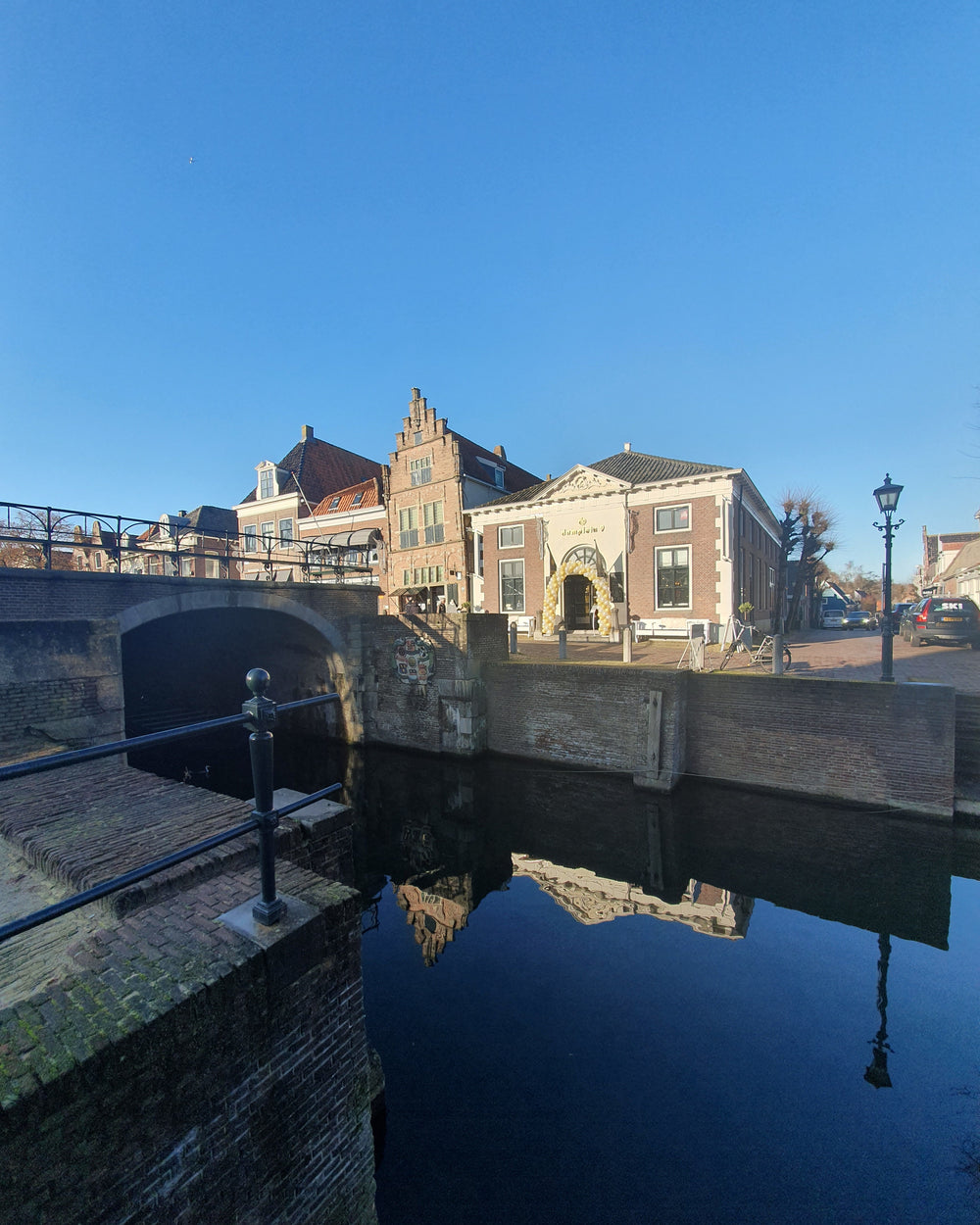
Lord Volture - It is nearly time for Christmas and then the New year... Where do you want to see Lord Volture LIVE in 2020? Let us know! Upcoming: 15.08.2020 – Zeeltje

PDF) Outer membrane permeabilization by the membrane attack complex sensitizes Gram-negative bacteria to antimicrobial proteins in serum and phagocytes

Unveiling the Inner Workings of Live Bacteria Using Super-Resolution Microscopy | Analytical Chemistry

Mismatch Repair Inhibits Homeologous Recombination via Coordinated Directional Unwinding of Trapped DNA Structures - ScienceDirect

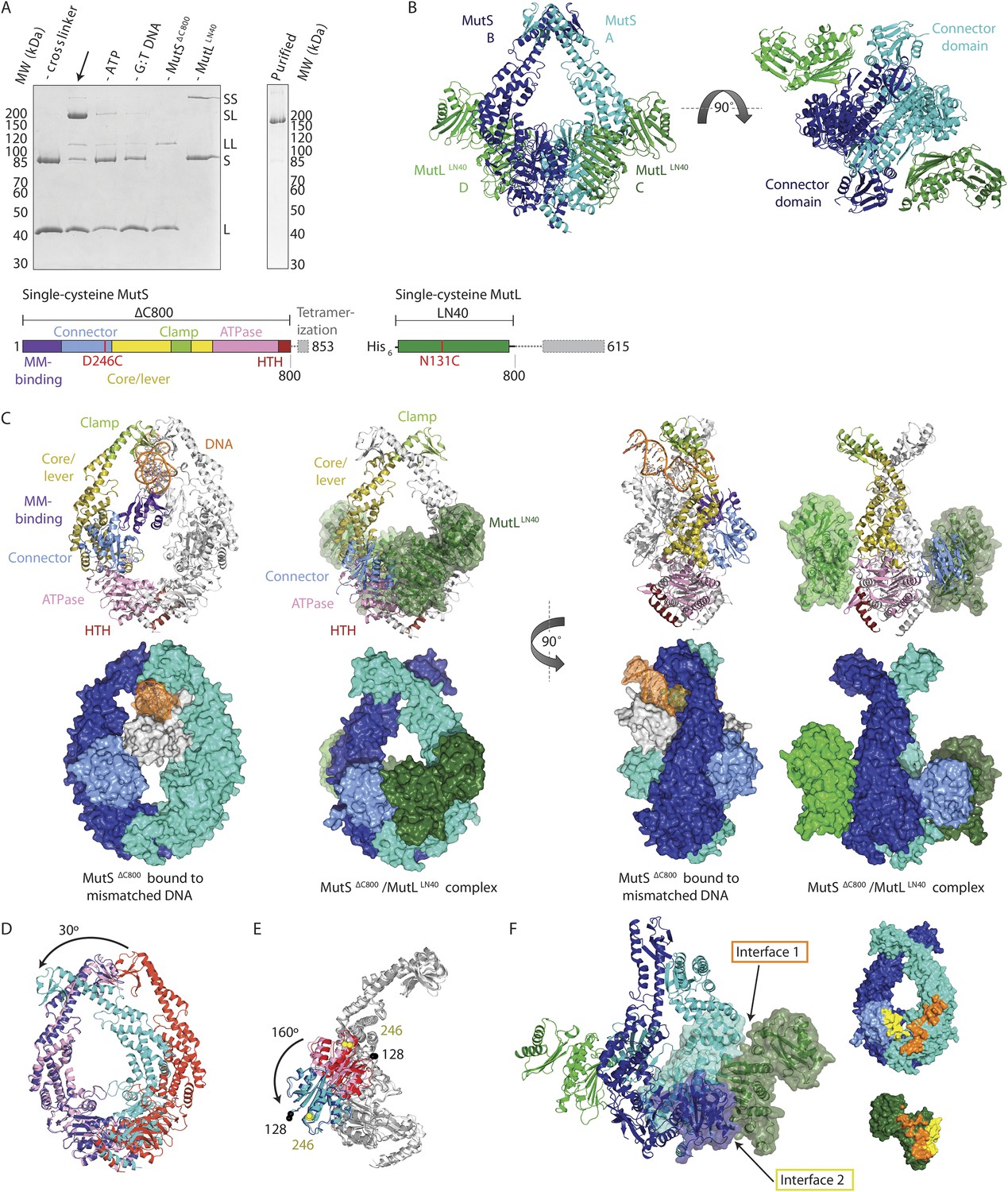
Cryo-EM structures reveal how ATP and DNA binding in MutS coordinate the sequential steps of DNA mismatch repair | bioRxiv

DNA mismatch and damage detection using a FRET-based assay for monitoring the loading of multiple MutS sliding clamps | bioRxiv

A spontaneous mutation in MutL‐Homolog 3 (HvMLH3) affects synapsis and crossover resolution in the barley desynaptic mutant des10 - Colas - 2016 - New Phytologist - Wiley Online Library

Cellular Recognition of Tri-/Di-palmitoylated Peptides Is Independent from a Domain Encompassing the N-terminal Seven Leucine-rich Repeat (LRR)/LRR-like Motifs of TLR2 - ScienceDirect

PDF) Using stable MutS dimers and tetramers to quantitatively analyze DNA mismatch recognition and sliding clamp formation

Dual daughter strand incision is processive and increases the efficiency of DNA mismatch repair. - Abstract - Europe PMC

PDF) Using stable MutS dimers and tetramers to quantitatively analyze DNA mismatch recognition and sliding clamp formation
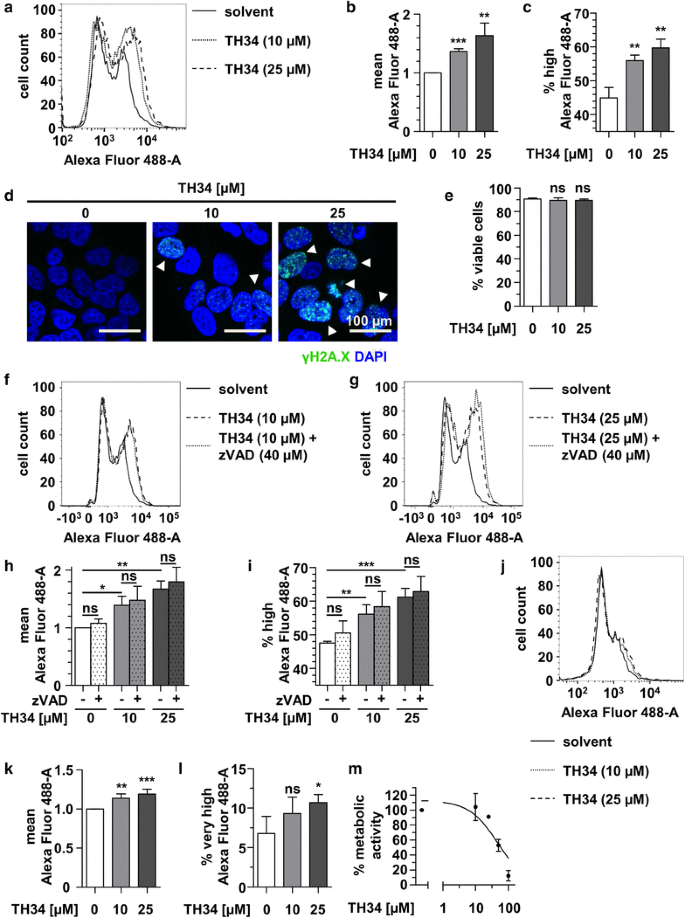
The HDAC6/8/10 inhibitor TH34 induces DNA damage-mediated cell death in human high-grade neuroblastoma cell lines | Archives of Toxicology

Cryogenic electron microscopy structures reveal how ATP and DNA binding in MutS coordinates sequential steps of DNA mismatch repair | Nature Structural & Molecular Biology

FIRRM/C1orf112 mediates resolution of homologous recombination intermediates in response to DNA interstrand crosslinks | Science Advances







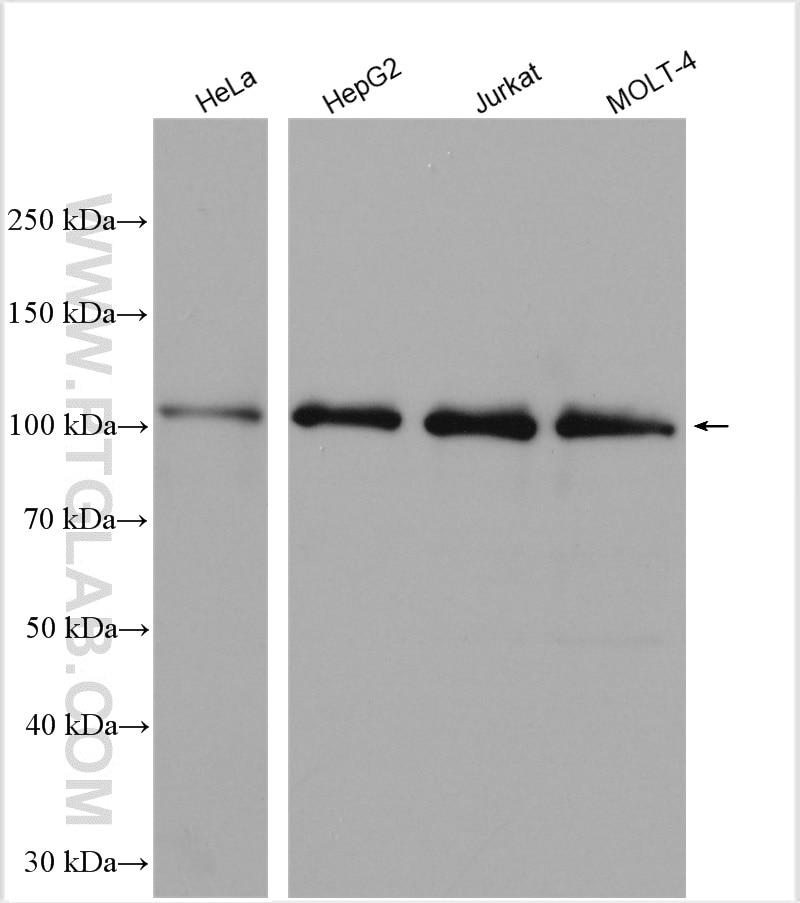
![Recombinant Anti-MLH1 antibody [EPR3894] KO Tested (ab92312) | Abcam Recombinant Anti-MLH1 antibody [EPR3894] KO Tested (ab92312) | Abcam](https://www.abcam.com/ps/products/92/ab92312/Images/ab92312-16-anti-mlh1-antibody-epr3894-hap-1-flow-cytometry.jpg)




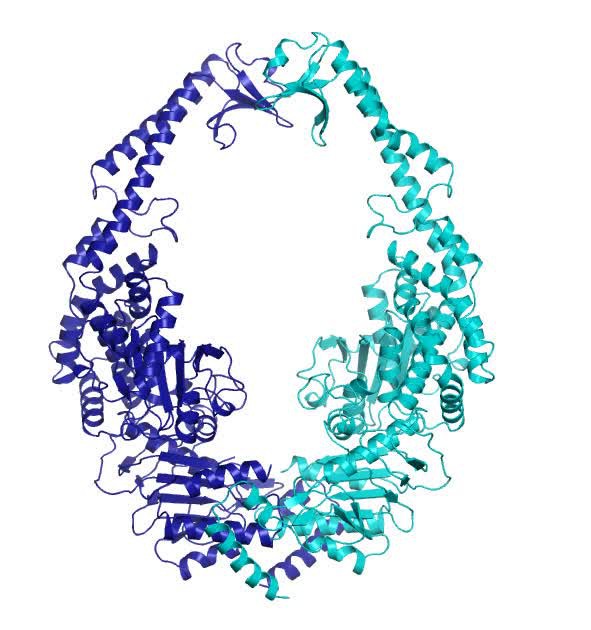


![Anti-MSH2 antibody [EPR21017-2] Recombinant (ab212188) | Abcam Anti-MSH2 antibody [EPR21017-2] Recombinant (ab212188) | Abcam](https://www.abcam.com/ps/products/212/ab212188/Images/ab212188-369004-anti-msh2-antibody-epr21017-2-msh2-ko-hap1-flow-cytometry-human.jpg)
![Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam](https://www.abcam.com/ps/products/199/ab199237/Images/ab199237-2-benefits-of-recombinant-antibodies.png)




![Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam Alexa Fluor® 488 Anti-MLH1 antibody [EPR3894] (ab199237) | Abcam](https://www.abcam.com/ps/products/199/ab199237/Images/ab199237-248777-anti-mlh1-antibody-epr3894-alexa-fluor-488-immunofluorescence.jpg)







